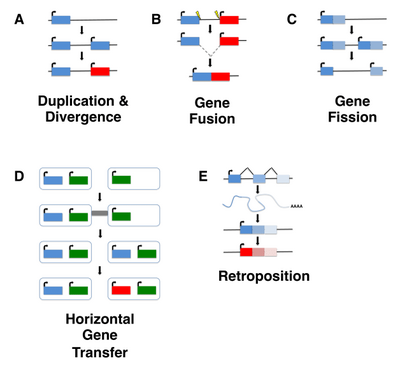
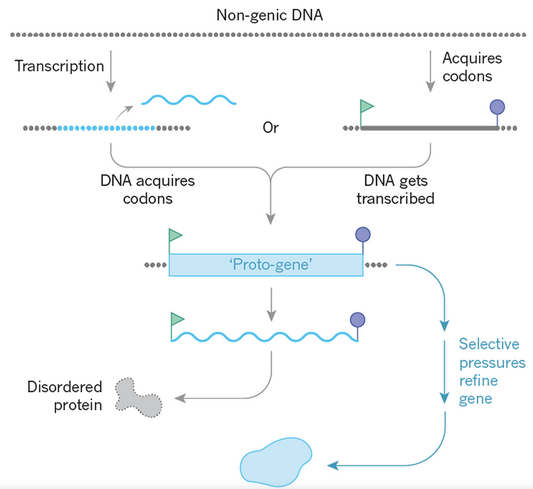
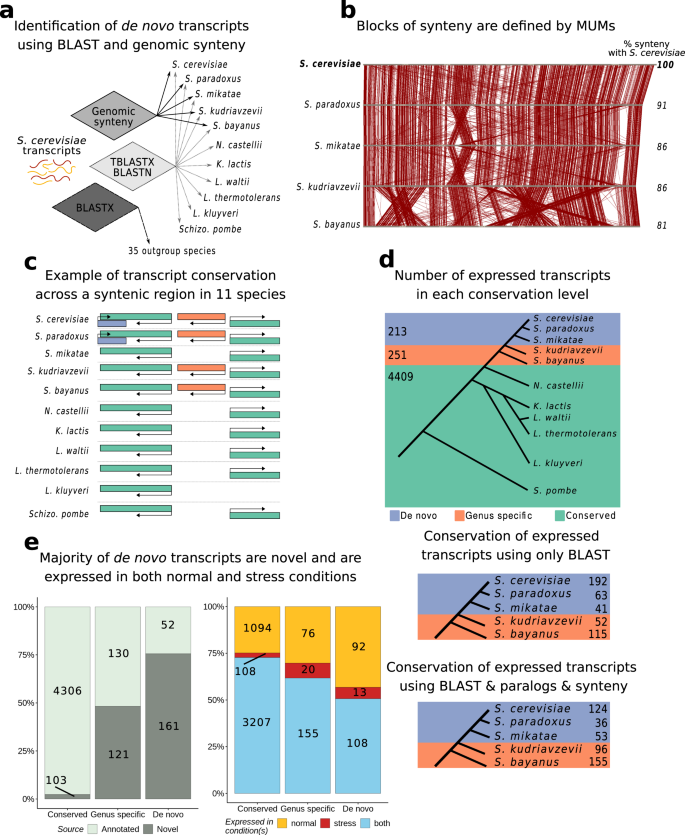
De Novo Transcriptome Sequencing of Ramonda serbica: Identification of the Candidate Genes Involved in the Desiccation Tolerance

De novo designed peptides for cellular delivery and subcellular localisation | Nature Chemical Biology

PDF) TransFlow: A modular framework for assembling and assessing accurate de novo transcriptomes in non-model organisms

Mustafa Kansiz en LinkedIn: Spatiotemporal Heterogeneity of De Novo Lipogenesis in Fixed and Living…
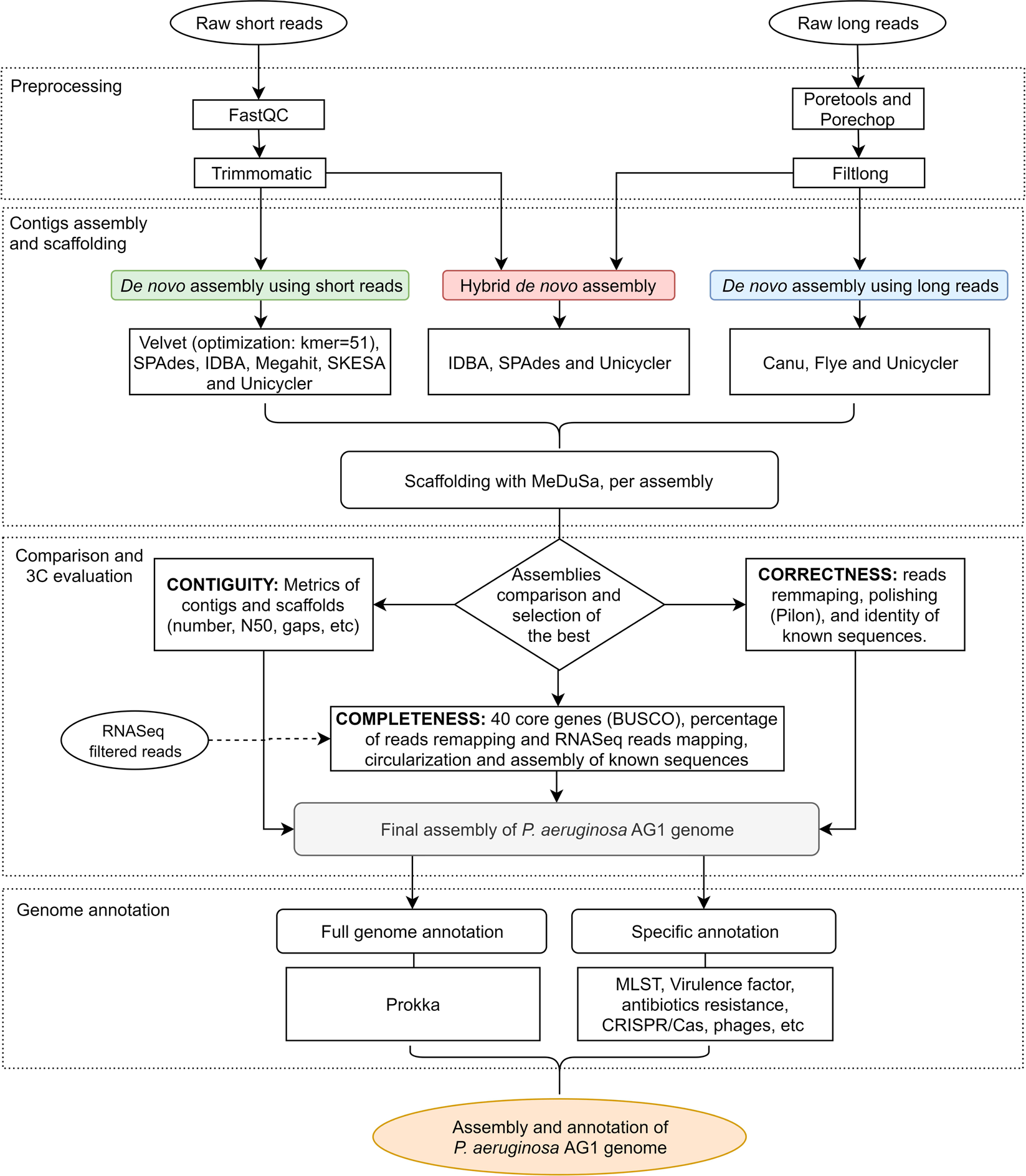
High quality 3C de novo assembly and annotation of a multidrug resistant ST-111 Pseudomonas aeruginosa genome: Benchmark of hybrid and non-hybrid assemblers | Scientific Reports